Validation
There are many ways to validate a model with scalecast and this notebook introduces them and overviews the differences between dynamic and non-dynamic tuning/testing, cross-validation, backtesting, and the eye test.
Download data: https://www.kaggle.com/robervalt/sunspots
See here for EDA on this dataset: https://scalecast-examples.readthedocs.io/en/latest/rnn/rnn.html
See here for documentation on cross validation: https://scalecast.readthedocs.io/en/latest/Forecaster/Forecaster.html#src.scalecast.Forecaster.Forecaster.cross_validate
[1]:
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from scalecast.Forecaster import Forecaster
import diagram_creator
[2]:
def prepare_fcst(f, test_length=0.1, fcst_length=120):
""" adds all variables and sets the test length/forecast length in the object
Args:
f (Forecaster): the Forecaster object.
test_length (int or float): the test length as a size or proportion.
fcst_length (int): the forecast horizon.
Returns:
(Forecaster) the processed object.
"""
f.generate_future_dates(fcst_length)
f.set_test_length(test_length)
f.set_validation_length(f.test_length)
f.eval_cis()
f.add_seasonal_regressors("month")
for i in np.arange(60, 289, 12): # 12-month cycles from 12 to 288 months
f.add_cycle(i)
f.add_ar_terms(120) # AR 1-120
f.add_AR_terms((20, 12)) # seasonal AR up to 20 years, spaced one year apart
f.add_seasonal_regressors("year")
#f.auto_Xvar_select(irr_cycles=[120],estimator='gbt')
return f
def export_results(f):
""" returns a dataframe with all model results given a Forecaster object.
Args:
f (Forecaster): the Forecaster object.
Returns:
(DataFrame) the dataframe with the pertinent results.
"""
results = f.export("model_summaries", determine_best_by="TestSetMAE")
return results[
[
"ModelNickname",
"TestSetRMSE",
"InSampleRMSE",
"ValidationMetric",
"ValidationMetricValue",
"HyperParams",
"TestSetLength",
"DynamicallyTested",
]
]
Load Forecaster Object
we choose 120 periods (10 years) for all validation and forecasting
10 years of observervations to tune model hyperparameters, 10 years to test, and a forecast horizon of 10 years
[3]:
df = pd.read_csv("Sunspots.csv", index_col=0, names=["Date", "Target"], header=0)
f = Forecaster(y=df["Target"], current_dates=df["Date"])
prepare_fcst(f)
[3]:
Forecaster(
DateStartActuals=1749-01-31T00:00:00.000000000
DateEndActuals=2021-01-31T00:00:00.000000000
Freq=M
N_actuals=3265
ForecastLength=120
Xvars=['month', 'cycle60sin', 'cycle60cos', 'cycle72sin', 'cycle72cos', 'cycle84sin', 'cycle84cos', 'cycle96sin', 'cycle96cos', 'cycle108sin', 'cycle108cos', 'cycle120sin', 'cycle120cos', 'cycle132sin', 'cycle132cos', 'cycle144sin', 'cycle144cos', 'cycle156sin', 'cycle156cos', 'cycle168sin', 'cycle168cos', 'cycle180sin', 'cycle180cos', 'cycle192sin', 'cycle192cos', 'cycle204sin', 'cycle204cos', 'cycle216sin', 'cycle216cos', 'cycle228sin', 'cycle228cos', 'cycle240sin', 'cycle240cos', 'cycle252sin', 'cycle252cos', 'cycle264sin', 'cycle264cos', 'cycle276sin', 'cycle276cos', 'cycle288sin', 'cycle288cos', 'AR1', 'AR2', 'AR3', 'AR4', 'AR5', 'AR6', 'AR7', 'AR8', 'AR9', 'AR10', 'AR11', 'AR12', 'AR13', 'AR14', 'AR15', 'AR16', 'AR17', 'AR18', 'AR19', 'AR20', 'AR21', 'AR22', 'AR23', 'AR24', 'AR25', 'AR26', 'AR27', 'AR28', 'AR29', 'AR30', 'AR31', 'AR32', 'AR33', 'AR34', 'AR35', 'AR36', 'AR37', 'AR38', 'AR39', 'AR40', 'AR41', 'AR42', 'AR43', 'AR44', 'AR45', 'AR46', 'AR47', 'AR48', 'AR49', 'AR50', 'AR51', 'AR52', 'AR53', 'AR54', 'AR55', 'AR56', 'AR57', 'AR58', 'AR59', 'AR60', 'AR61', 'AR62', 'AR63', 'AR64', 'AR65', 'AR66', 'AR67', 'AR68', 'AR69', 'AR70', 'AR71', 'AR72', 'AR73', 'AR74', 'AR75', 'AR76', 'AR77', 'AR78', 'AR79', 'AR80', 'AR81', 'AR82', 'AR83', 'AR84', 'AR85', 'AR86', 'AR87', 'AR88', 'AR89', 'AR90', 'AR91', 'AR92', 'AR93', 'AR94', 'AR95', 'AR96', 'AR97', 'AR98', 'AR99', 'AR100', 'AR101', 'AR102', 'AR103', 'AR104', 'AR105', 'AR106', 'AR107', 'AR108', 'AR109', 'AR110', 'AR111', 'AR112', 'AR113', 'AR114', 'AR115', 'AR116', 'AR117', 'AR118', 'AR119', 'AR120', 'AR132', 'AR144', 'AR156', 'AR168', 'AR180', 'AR192', 'AR204', 'AR216', 'AR228', 'AR240', 'year']
TestLength=326
ValidationMetric=rmse
ForecastsEvaluated=[]
CILevel=0.95
CurrentEstimator=mlr
GridsFile=Grids
)
[4]:
f.set_estimator('gbt')
In the feature_selection notebook, gbt was chosen as the best model class out of several tried. We will show all examples with this estimator.
Default Model Parameters
one with dynamic testing
one with non-dynamic testing
the difference can be expressed by taking the case of a simple autoregressive model, such that:
Non-Dynamic Autoregressive Predictions
Over an indefinite forecast horizon, the above equation would only work if you knew the value for \(x_{t-1}\). Going more than one period into the future, you would stop knowing what that value is. In a test-set of data, of course, you do know all values into the forecast horizon, but to be more realistic, you could write an equation for a two-step forecast like this:
Dynamic Autoregressive Predictions
Using these two equations, which scalecast refers to as dynamic forecasting, you could evaluate any forecast horizon by plugging in predicted values for \(x_{t-1}\) or \({x_t}\) over periods in which you did not know it. This is default behavior for all models tested through scalecast, but it is not default for tuning models. We will explore dynamic tuning soon. First, let’s see in practical terms the difference between non-dynamic and dynamic testing.
[5]:
f.manual_forecast(call_me="gbt_default_non-dynamic", dynamic_testing=False)
f.manual_forecast(call_me="gbt_default_dynamic") # default is dynamic testing
[6]:
f.plot_test_set(
models=[
"gbt_default_non-dynamic",
"gbt_default_dynamic"
],
include_train=False,
)
plt.show()
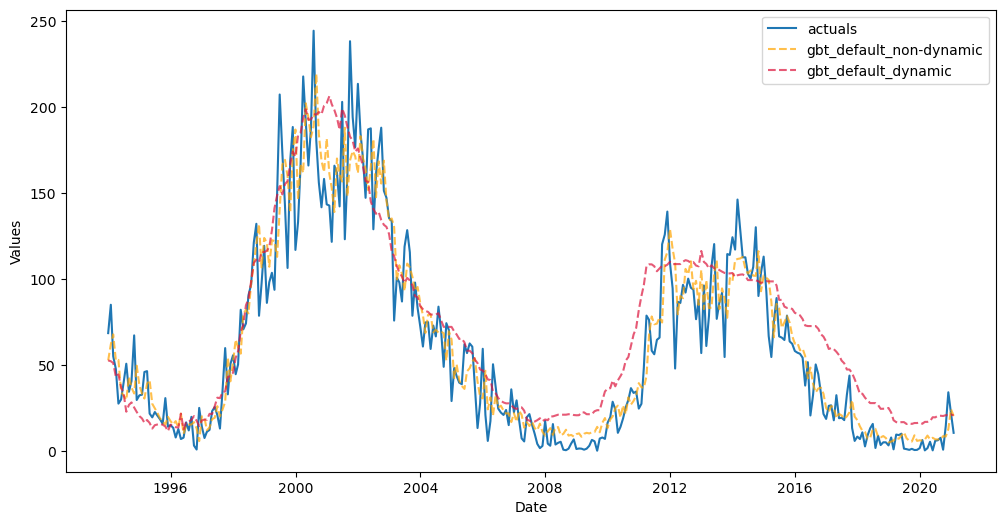
It appears that the non-dynamically tested model performed significantly better than the other, but looks can be deceiving. In essence, the non-dynamic model was only tested for its ability to perform 326 one-step forecasts and its final metric is an average of these one-step forecasts. It could be a good idea to set dynamic_testing=False if you want to speed up the testing process or if you only care about how your model would perform one step into the future. But to report the test-set
metric from this model as if it could be expected to do that well for the full 326 periods into the future is misleading. The other model that was dynamically tested can be more realistically trusted in that regard.
[7]:
export_results(f)
[7]:
| ModelNickname | TestSetRMSE | InSampleRMSE | ValidationMetric | ValidationMetricValue | HyperParams | TestSetLength | DynamicallyTested | |
|---|---|---|---|---|---|---|---|---|
| 0 | gbt_default_non-dynamic | 18.723179 | 18.841544 | NaN | NaN | {} | 326 | False |
| 1 | gbt_default_dynamic | 24.456923 | 18.841544 | NaN | NaN | {} | 326 | True |
Tune the model to find optimal hyperparameters
Create a validation grid
Try three strategies to tune the parameters:
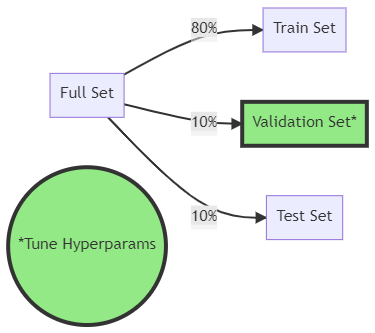
Train/validation/test split
Hyperparameters are tried on the validation set
Train/test split with 5-fold time-series cross-validation on training set
Training data split 5 times into train/validations set
Models trained on training set only
Validated out-of-sample
All data available before each validation split sequentially used to train the model
Train/test split with 5-fold time-series rolling cross-validation on training set
Rolling is different in that each train/validation split is the same size
[8]:
grid = {
"max_depth": [2, 3, 5],
"max_features": ["sqrt", "auto"],
"subsample": [0.8, 0.9, 1],
}
[9]:
f.ingest_grid(grid)
Train/Validation/Test Split
The data’s sequence is always maintained in time-series splits with scalecast
[10]:
f.tune(dynamic_tuning=True)
f.auto_forecast(
call_me="gbt_tuned"
) # automatically uses optimal paramaeters suggested from the tuning process
[11]:
f.export_validation_grid("gbt_tuned").sort_values('AverageMetric').head(10)
[11]:
| max_depth | max_features | subsample | Fold0Metric | AverageMetric | MetricEvaluated | |
|---|---|---|---|---|---|---|
| 7 | 3 | sqrt | 0.9 | 43.623057 | 43.623057 | rmse |
| 12 | 5 | sqrt | 0.8 | 43.667072 | 43.667072 | rmse |
| 6 | 3 | sqrt | 0.8 | 44.015361 | 44.015361 | rmse |
| 8 | 3 | sqrt | 1 | 44.807485 | 44.807485 | rmse |
| 0 | 2 | sqrt | 0.8 | 46.093143 | 46.093143 | rmse |
| 2 | 2 | sqrt | 1 | 53.206496 | 53.206496 | rmse |
| 14 | 5 | sqrt | 1 | 60.587243 | 60.587243 | rmse |
| 10 | 3 | auto | 0.9 | 65.424393 | 65.424393 | rmse |
| 15 | 5 | auto | 0.8 | 66.530315 | 66.530315 | rmse |
| 13 | 5 | sqrt | 0.9 | 67.459775 | 67.459775 | rmse |
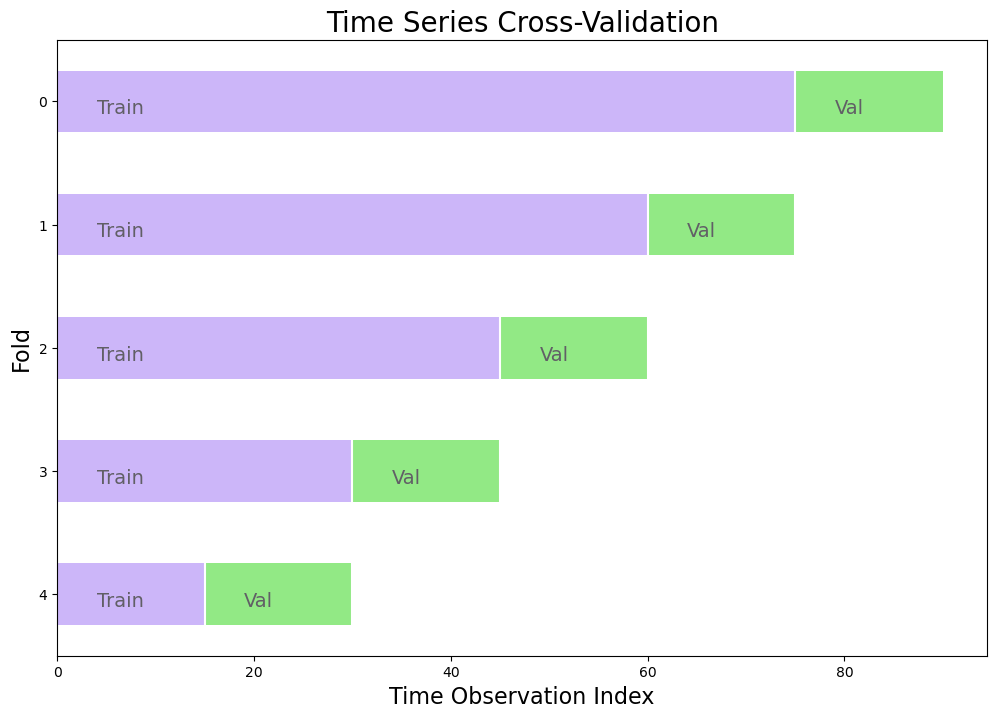
5-Fold Time Series Cross Validation
Split training set into k (5) folds
Each validation set is the same size and determined such that:
val_size = n_obs // (folds + 1)The last training set will be the same size or almost the same size as the validation sets
Model trained and re-trained with all data that came before each validation slice
Each fold tested out of sample on its validation set
Final error is an average of the out-of-sample error obtained from each fold
The chosen hyperparameters are determined by which final error was minimized
Below is an example with a dataset sized 100 observations and in which 10 observations are held out for testing and the remaining 90 observations are used as the training set:
[12]:
diagram_creator.create_cv()
The final error, E, can be expressed as an average of the error from each fold i:
[13]:
f.cross_validate(
k=5,
dynamic_tuning=True,
test_length = None, # default so that last test and train sets are same size (or close to the same)
train_length = None, # default uses all observations before each test set
space_between_sets = None, # default adds a length equal to the test set between consecutive sets
verbose = True, # print out info about each fold
)
f.auto_forecast(call_me="gbt_cv")
Num hyperparams to try for the gbt model: 18.
Fold 0: Train size: 2450 (1749-01-31 00:00:00 - 1953-02-28 00:00:00). Test Size: 489 (1953-03-31 00:00:00 - 1993-11-30 00:00:00).
Fold 1: Train size: 1961 (1749-01-31 00:00:00 - 1912-05-31 00:00:00). Test Size: 489 (1912-06-30 00:00:00 - 1953-02-28 00:00:00).
Fold 2: Train size: 1472 (1749-01-31 00:00:00 - 1871-08-31 00:00:00). Test Size: 489 (1871-09-30 00:00:00 - 1912-05-31 00:00:00).
Fold 3: Train size: 983 (1749-01-31 00:00:00 - 1830-11-30 00:00:00). Test Size: 489 (1830-12-31 00:00:00 - 1871-08-31 00:00:00).
Fold 4: Train size: 494 (1749-01-31 00:00:00 - 1790-02-28 00:00:00). Test Size: 489 (1790-03-31 00:00:00 - 1830-11-30 00:00:00).
Chosen paramaters: {'max_depth': 5, 'max_features': 'sqrt', 'subsample': 1}.
[14]:
f.export_validation_grid("gbt_cv").sort_values("AverageMetric").head(10)
[14]:
| max_depth | max_features | subsample | Fold0Metric | Fold1Metric | Fold2Metric | Fold3Metric | Fold4Metric | AverageMetric | MetricEvaluated | |
|---|---|---|---|---|---|---|---|---|---|---|
| 14 | 5 | sqrt | 1 | 75.881634 | 85.585377 | 79.162305 | 83.264294 | 96.912215 | 84.161165 | rmse |
| 12 | 5 | sqrt | 0.8 | 90.328729 | 96.703967 | 48.606034 | 80.703070 | 111.586242 | 85.585609 | rmse |
| 13 | 5 | sqrt | 0.9 | 108.327532 | 95.594713 | 61.082475 | 61.798101 | 108.437289 | 87.048022 | rmse |
| 16 | 5 | auto | 0.9 | 89.047882 | 92.811981 | 74.332318 | 86.321627 | 112.510827 | 91.004927 | rmse |
| 0 | 2 | sqrt | 0.8 | 105.541098 | 78.227464 | 103.386493 | 77.997451 | 115.573720 | 96.145245 | rmse |
| 8 | 3 | sqrt | 1 | 64.668019 | 119.509341 | 93.877028 | 93.740770 | 114.533147 | 97.265661 | rmse |
| 7 | 3 | sqrt | 0.9 | 75.061438 | 85.898970 | 122.575339 | 91.157722 | 119.005642 | 98.739822 | rmse |
| 2 | 2 | sqrt | 1 | 59.907395 | 88.529044 | 113.535870 | 118.721240 | 117.856393 | 99.709989 | rmse |
| 17 | 5 | auto | 1 | 111.676983 | 96.506098 | 70.688360 | 102.882864 | 118.365703 | 100.024002 | rmse |
| 6 | 3 | sqrt | 0.8 | 93.640946 | 85.464725 | 100.451257 | 109.813079 | 111.415691 | 100.157140 | rmse |
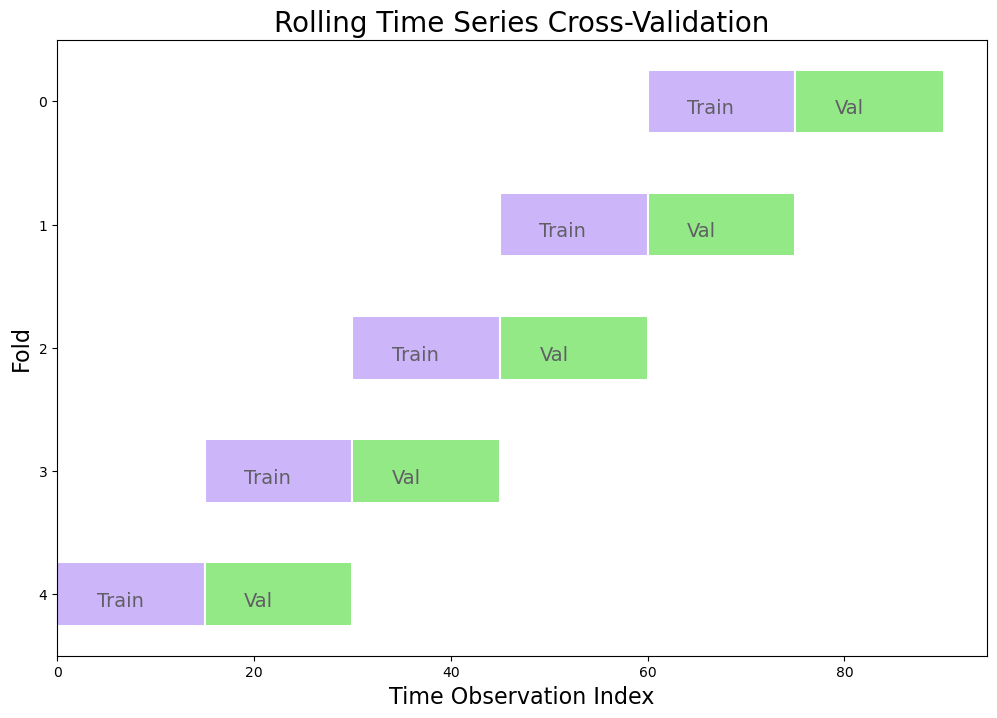
5-Fold Rolling Time Series Cross Validation
Split training set into k (5) folds
Each validation set is the same size
Each training set is also the same size as each validation set
Each fold tested out of sample
Final error is an average of the out-of-sample error obtained from each folds
The chosen hyperparameters are determined by which final error was minimized
Below is an example with a dataset sized 100 observation and in which 10 observations are held out for testing and the remaining 90 observations are used as the training set:
[15]:
diagram_creator.create_rolling_cv()
[16]:
f.cross_validate(
k=5,
rolling=True,
dynamic_tuning=True,
test_length = None, # with rolling = True, makes all train and test sets the same size
train_length = None, # with rolling = True, makes all train and test sets the same size
space_between_sets = None, # default adds a length equal to the test set between consecutive sets
verbose = True, # print out info about each fold
)
f.auto_forecast(call_me="gbt_rolling_cv")
Num hyperparams to try for the gbt model: 18.
Fold 0: Train size: 489 (1912-06-30 00:00:00 - 1953-02-28 00:00:00). Test Size: 489 (1953-03-31 00:00:00 - 1993-11-30 00:00:00).
Fold 1: Train size: 489 (1871-09-30 00:00:00 - 1912-05-31 00:00:00). Test Size: 489 (1912-06-30 00:00:00 - 1953-02-28 00:00:00).
Fold 2: Train size: 489 (1830-12-31 00:00:00 - 1871-08-31 00:00:00). Test Size: 489 (1871-09-30 00:00:00 - 1912-05-31 00:00:00).
Fold 3: Train size: 489 (1790-03-31 00:00:00 - 1830-11-30 00:00:00). Test Size: 489 (1830-12-31 00:00:00 - 1871-08-31 00:00:00).
Fold 4: Train size: 489 (1749-06-30 00:00:00 - 1790-02-28 00:00:00). Test Size: 489 (1790-03-31 00:00:00 - 1830-11-30 00:00:00).
Chosen paramaters: {'max_depth': 5, 'max_features': 'sqrt', 'subsample': 0.9}.
[17]:
f.export_validation_grid("gbt_rolling_cv").sort_values("AverageMetric").head(10)
[17]:
| max_depth | max_features | subsample | Fold0Metric | Fold1Metric | Fold2Metric | Fold3Metric | Fold4Metric | AverageMetric | MetricEvaluated | |
|---|---|---|---|---|---|---|---|---|---|---|
| 13 | 5 | sqrt | 0.9 | 52.067612 | 75.101589 | 47.665446 | 73.458303 | 110.673685 | 71.793327 | rmse |
| 11 | 3 | auto | 1 | 61.780860 | 70.554830 | 48.914548 | 74.467054 | 110.351684 | 73.213795 | rmse |
| 9 | 3 | auto | 0.8 | 59.526670 | 76.807231 | 48.096324 | 74.276742 | 110.628222 | 73.867038 | rmse |
| 17 | 5 | auto | 1 | 68.884924 | 63.475008 | 52.178487 | 73.895434 | 111.100209 | 73.906812 | rmse |
| 7 | 3 | sqrt | 0.9 | 44.798329 | 69.131312 | 54.226665 | 93.287371 | 115.134538 | 75.315643 | rmse |
| 8 | 3 | sqrt | 1 | 56.409051 | 84.081666 | 49.339527 | 75.453036 | 111.791536 | 75.414963 | rmse |
| 3 | 2 | auto | 0.8 | 61.498220 | 73.383101 | 48.898466 | 78.773421 | 115.272207 | 75.565083 | rmse |
| 12 | 5 | sqrt | 0.8 | 63.732031 | 78.140692 | 47.053796 | 73.473322 | 115.558499 | 75.591668 | rmse |
| 5 | 2 | auto | 1 | 69.075181 | 67.852121 | 53.383812 | 75.150129 | 115.413148 | 76.174878 | rmse |
| 14 | 5 | sqrt | 1 | 48.860162 | 84.807936 | 49.921226 | 89.144401 | 110.995682 | 76.745881 | rmse |
View results
[18]:
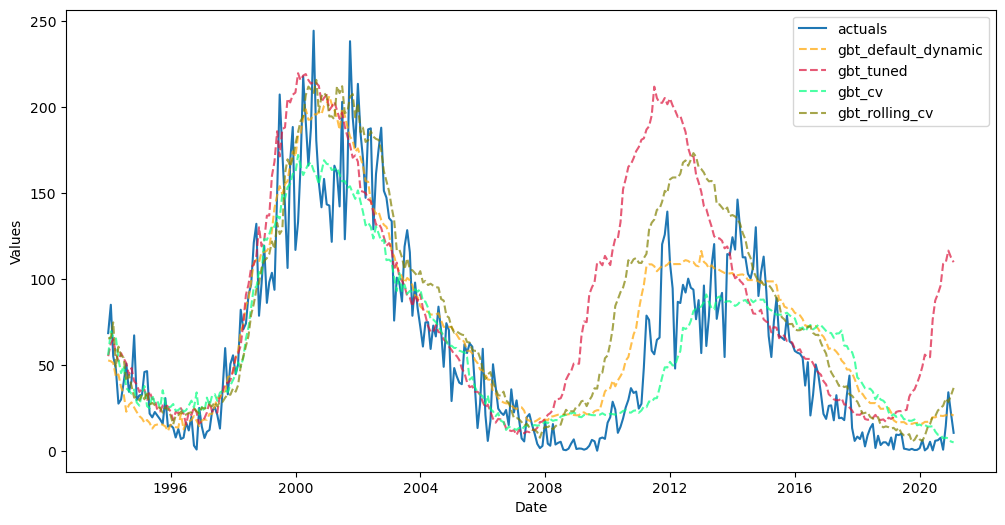
f.plot_test_set(
models=[
"gbt_default_dynamic",
"gbt_tuned",
"gbt_cv",
"gbt_rolling_cv",
],
include_train=False,
)
plt.show()
[19]:
pd.set_option('max_colwidth', 60)
export_results(f)
[19]:
| ModelNickname | TestSetRMSE | InSampleRMSE | ValidationMetric | ValidationMetricValue | HyperParams | TestSetLength | DynamicallyTested | |
|---|---|---|---|---|---|---|---|---|
| 0 | gbt_default_non-dynamic | 18.723179 | 18.841544 | NaN | NaN | {} | 326 | False |
| 1 | gbt_default_dynamic | 24.456923 | 18.841544 | NaN | NaN | {} | 326 | True |
| 2 | gbt_cv | 25.604943 | 13.063379 | rmse | 84.161165 | {'max_depth': 5, 'max_features': 'sqrt', 'subsample': 1} | 326 | True |
| 3 | gbt_rolling_cv | 34.248137 | 12.789779 | rmse | 71.793327 | {'max_depth': 5, 'max_features': 'sqrt', 'subsample': 0.9} | 326 | True |
| 4 | gbt_tuned | 50.777958 | 20.342340 | rmse | 43.623057 | {'max_depth': 3, 'max_features': 'sqrt', 'subsample': 0.9} | 326 | True |
Backtest models
Backtesting is a process in which the final chosen model is re-validated by seeing its average performance on the last x-number of forecast horizons available in the data
With scalecast, pipeline objects can be built and backtest all applied models to see the best one over several forecast horizons
See the documentation for more information
[20]:
from scalecast.Pipeline import Pipeline
from scalecast.util import backtest_metrics
[21]:
def forecaster(f):
f.set_estimator('gbt')
f.set_validation_length(int(len(f.y)*.1)) # 10% val length each time
f.add_seasonal_regressors("month")
for i in np.arange(60, 289, 12): # 12-month cycles from 12 to 288 months
f.add_cycle(i)
f.add_ar_terms(120) # AR 1-120
f.add_AR_terms((20, 12)) # seasonal AR up to 20 years, spaced one year apart
f.add_seasonal_regressors("year")
f.manual_forecast(call_me='gbt_default_dynamic')
f.ingest_grid(grid)
f.tune(dynamic_tuning = True)
f.auto_forecast(call_me='gbt_tuned')
f.cross_validate(dynamic_tuning = True)
f.auto_forecast(call_me='gbt_cv')
f.cross_validate(dynamic_tuning = True, rolling = True)
f.auto_forecast(call_me='gbt_rolling_cv')
[22]:
pipeline = Pipeline(steps = [('Forecast',forecaster)])
[23]:
backtest_results = pipeline.backtest(
f,
test_length = 0,
fcst_length = 120,
jump_back = 120, # place 120 obs between each training set
cis = False,
verbose = True,
)
[24]:
bm = backtest_metrics(backtest_results,mets=['rmse','mae','r2'])
bm
[24]:
| Iter0 | Iter1 | Iter2 | Iter3 | Iter4 | Average | ||
|---|---|---|---|---|---|---|---|
| Model | Metric | ||||||
| gbt_default_dynamic | rmse | 29.787489 | 32.631984 | 40.218024 | 57.655588 | 88.550393 | 49.768696 |
| mae | 24.725663 | 25.120353 | 32.70369 | 45.658795 | 56.387906 | 36.919281 | |
| r2 | 0.499971 | 0.718315 | 0.652106 | 0.466619 | -0.415585 | 0.384285 | |
| gbt_tuned | rmse | 21.793363 | 38.201181 | 34.145031 | 55.637363 | 99.862839 | 49.927956 |
| mae | 17.232072 | 29.314773 | 26.055781 | 42.62898 | 63.122565 | 35.670834 | |
| r2 | 0.732345 | 0.613962 | 0.749239 | 0.503307 | -0.800374 | 0.359696 | |
| gbt_cv | rmse | 27.290614 | 30.871908 | 38.859572 | 55.141773 | 35.648112 | 37.562396 |
| mae | 21.956079 | 24.026144 | 30.078019 | 42.157045 | 27.85357 | 29.214172 | |
| r2 | 0.580286 | 0.747883 | 0.675211 | 0.512116 | 0.770582 | 0.657215 | |
| gbt_rolling_cv | rmse | 26.080319 | 31.8572 | 34.485633 | 43.732603 | 38.312404 | 34.893632 |
| mae | 21.003643 | 24.701014 | 27.536833 | 37.057153 | 28.484026 | 27.756534 | |
| r2 | 0.616687 | 0.731533 | 0.744211 | 0.693122 | 0.735007 | 0.704112 |
[25]:
bmr = bm.reset_index()
[26]:
bmrrmse = bmr.loc[bmr['Metric'] == 'rmse'].sort_values('Average')
best_model = bmrrmse.iloc[0,0]
best_model
[26]:
'gbt_rolling_cv'
[27]:
rmse = bmr.loc[
(bmr['Model'] == best_model) & (bmr['Metric'] == 'rmse'),
'Average',
].values[0]
mae = bmr.loc[
(bmr['Model'] == best_model) & (bmr['Metric'] == 'mae'),
'Average',
].values[0]
r2 = bmr.loc[
(bmr['Model'] == best_model) & (bmr['Metric'] == 'r2'),
'Average',
].values[0]
[28]:
print(
f"The best model, according to the RMSE over 5 backtest iterations was {best_model}."
" On average, we can expect this model to have an RMSE of"
f" {rmse:.2f}, MAE of {mae:.2f}, and R2 of {r2:.2%} over a full 120-period"
" forecast window."
)
The best model, according to the RMSE over 5 backtest iterations was gbt_rolling_cv. On average, we can expect this model to have an RMSE of 34.89, MAE of 27.76, and R2 of 70.41% over a full 120-period forecast window.
An extension of this analysis could be to choose regressors more carefully (see here) and to use more complex models (see here).
The Eye Test
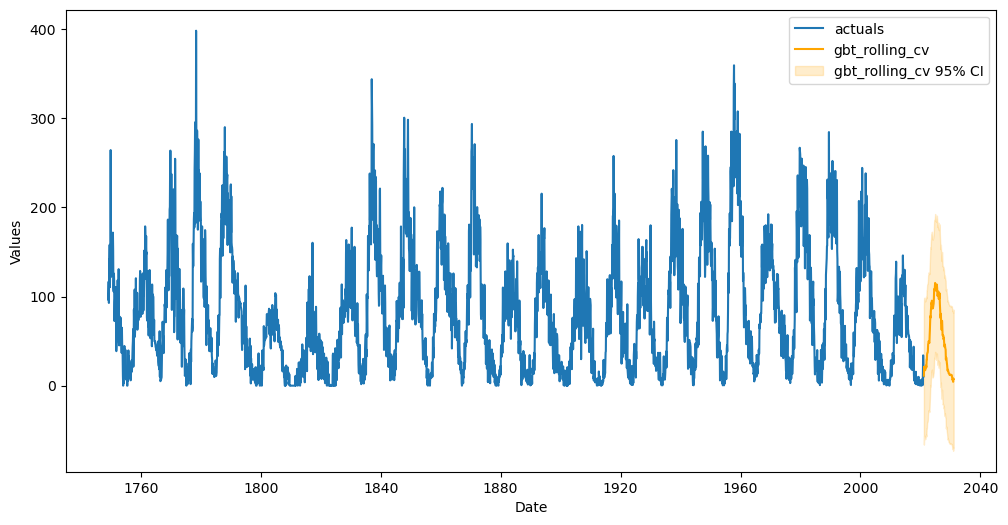
In addition to all the objective validation performed in this notebook, one of the most important questions to ask is if the forecast looks reasonable. Does it pass the common sense test? Below, we plot the 120-forecast period horizon from the best model.
[29]:
f.plot(models=best_model, ci = True)
plt.show()
[ ]: